Extracting Intake Forms with BAML and CocoIndex
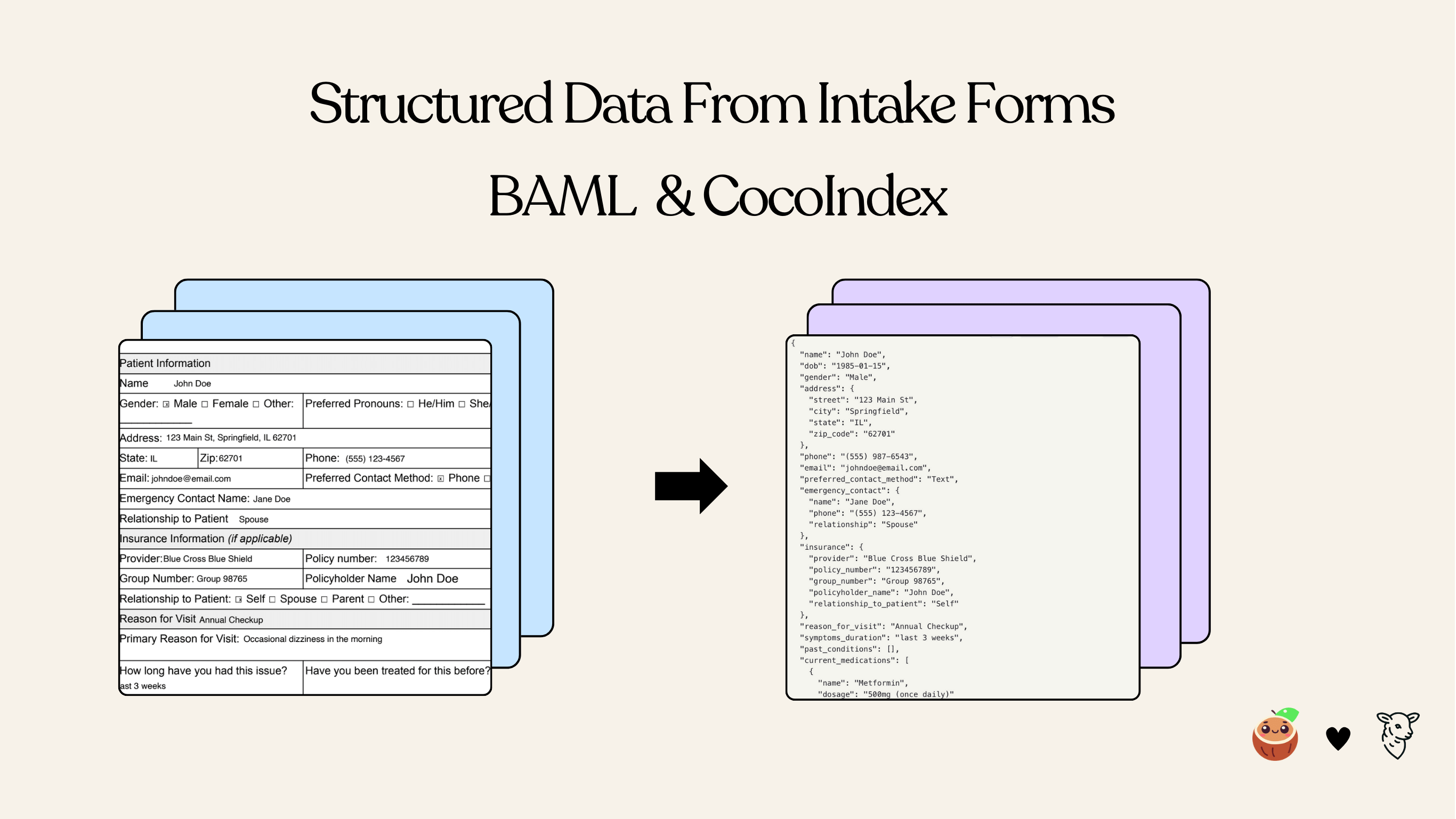

Overview
This tutorial shows how to use BAML together with CocoIndex to build a data pipeline that extracts structured patient information from PDF intake forms. The BAML definitions describe the desired output schema and prompt logic, while CocoIndex orchestrates file input, transformation, and incremental indexing.
The extraction quality is highly dependent on the OCR quality. You can use CocoIndex with any commercial parser or open source ones that is tailored for your domain for better results. For example, Document AI from Google Cloud and more.
Flow Overview
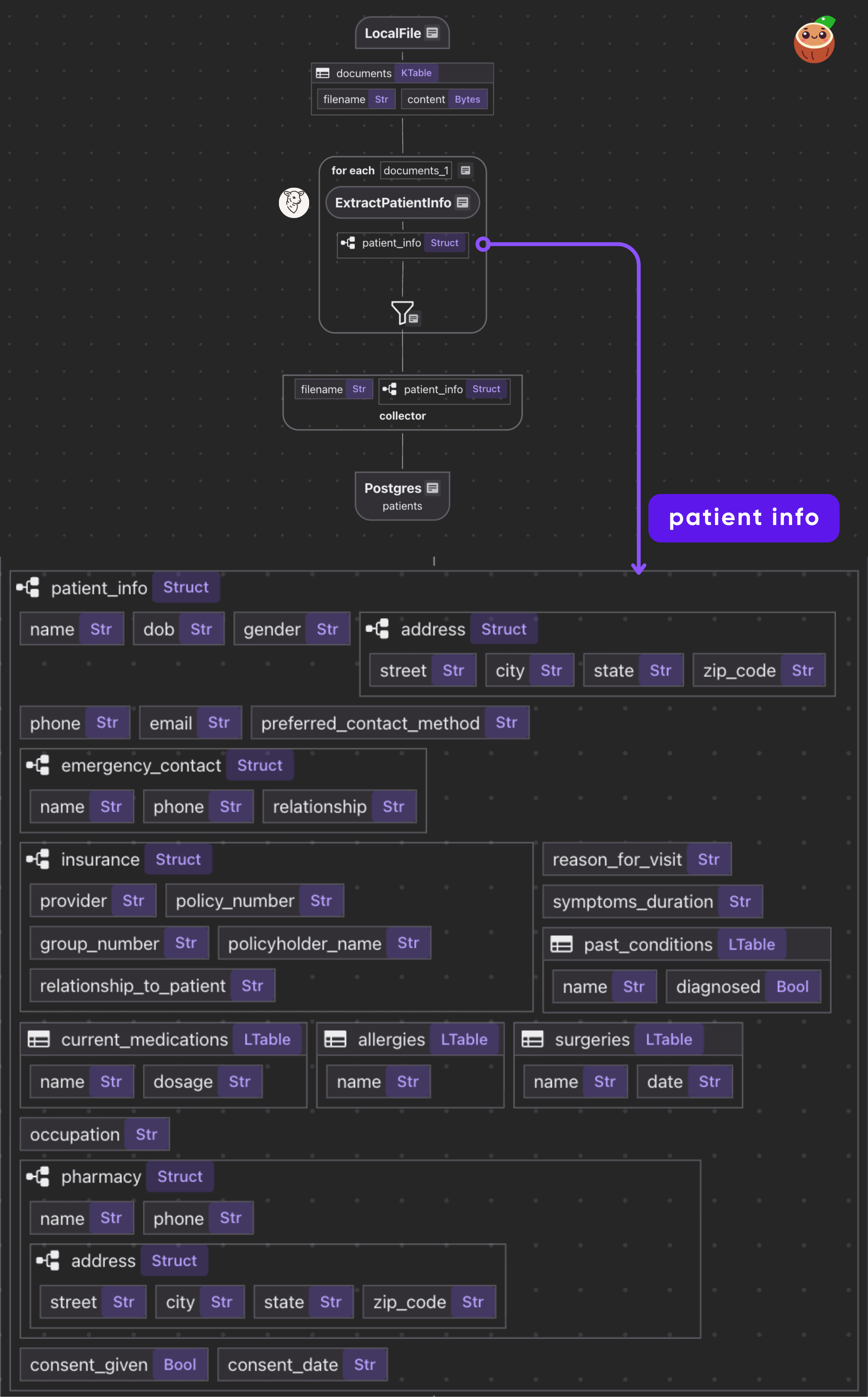
The flow itself is fairly simple.
- Read PDF files from a directory.
- For each file, call the BAML function to get a structured
Patient. - Collect results and export to Postgres.
Setup
-
Install Postgres if you don't have one.
-
Install dependencies
pip install -U cocoindex baml-py -
Create a
.envfile. You can copy it from.env.examplefirst:cp .env.example .envThen edit the file to fill in your
GEMINI_API_KEY.
Structured Extraction Component with BAML
Create a baml_src/ directory for your BAML definitions. We’ll define a schema for patient intake data (nested classes) and a function that prompts Gemini to extract those fields from a PDF. Save this as baml_src/patient.baml
Define Patient Schema
Classes: We defined Pydantic-style classes (Contact, Address, Insurance, etc.) to match the FHIR-inspired patient schema. These become typed output models. Required fields are non-nullable; optional fields use ?.
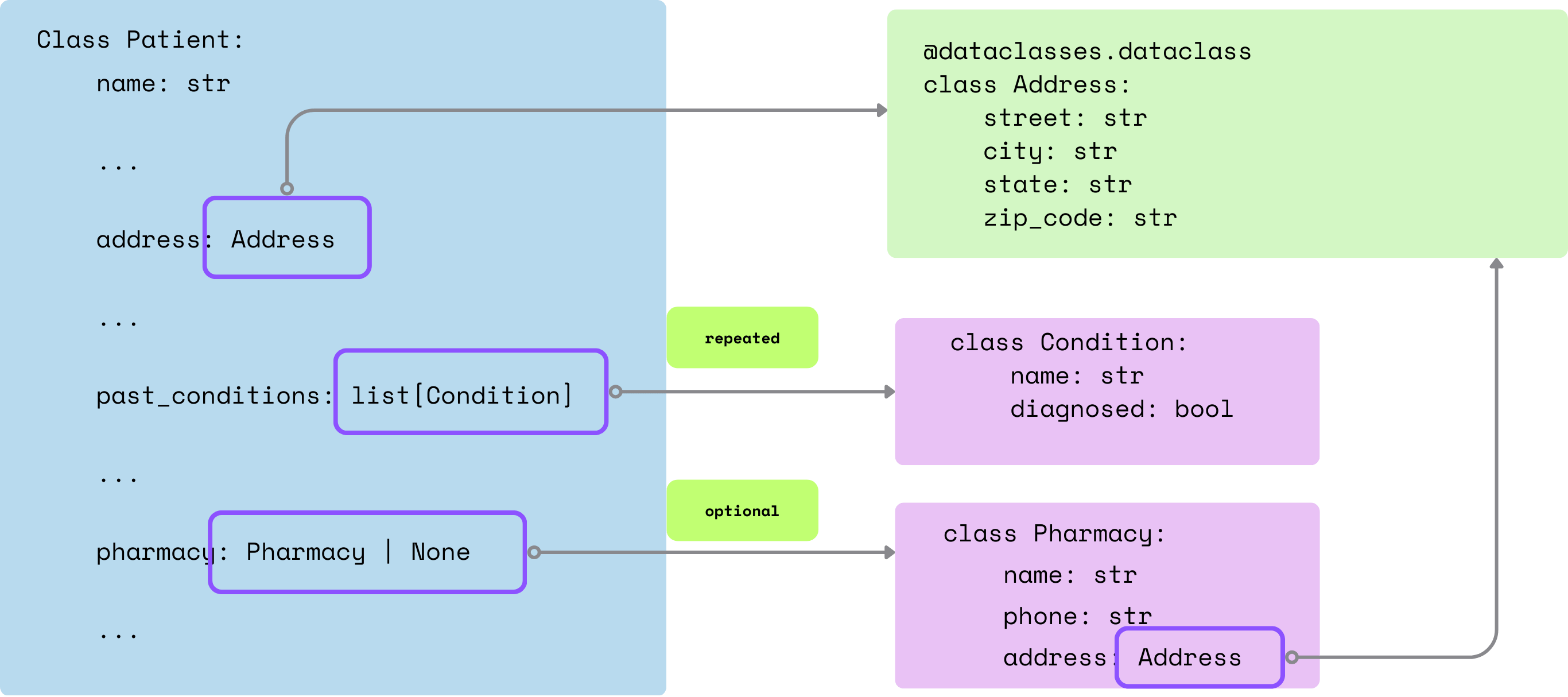
class Contact {
name string
phone string
relationship string
}
class Address {
street string
city string
state string
zip_code string
}
class Pharmacy {
name string
phone string
address Address
}
class Insurance {
provider string
policy_number string
group_number string?
policyholder_name string
relationship_to_patient string
}
class Condition {
name string
diagnosed bool
}
class Medication {
name string
dosage string
}
class Allergy {
name string
}
class Surgery {
name string
date string
}
class Patient {
name string
dob string
gender string
address Address
phone string
email string
preferred_contact_method string
emergency_contact Contact
insurance Insurance?
reason_for_visit string
symptoms_duration string
past_conditions Condition[]
current_medications Medication[]
allergies Allergy[]
surgeries Surgery[]
occupation string?
pharmacy Pharmacy?
consent_given bool
consent_date string?
}
Define the BAML function to extract patient info from a PDF
function ExtractPatientInfo(intake_form: pdf) -> Patient {
client Gemini
prompt #"
Extract all patient information from the following intake form document.
Please be thorough and extract all available information accurately.
{{ _.role("user") }}
{{ intake_form }}
Fill in with "N/A" for required fields if the information is not available.
{{ ctx.output_format }}
"#
}
We specify client Gemini and a prompt template. The special variable {{ intake_form }} injects the PDF, and {{ ctx.output_format }} tells BAML to expect the structured format defined by the return type. The prompt explicitly asks Gemini to extract all fields, filling “N/A” if missing.
role("user") Matters in BAML ExtractionIn our BAML example above, there's a subtle but crucial line: {{ _.role("user") }} is added at the start of the prompt.
This ensures the PDF content is explicitly included as part of the user message*, rather than the system prompt.
For OpenAI models, if the PDF is not in the user role, the model doesn't see the file content — so extractions will fail or return empty fields. This can easily trip you up.
Configure the LLM client to use Google’s Gemini model
client<llm> Gemini {
provider google-ai
options {
model gemini-2.5-flash
api_key env.GEMINI_API_KEY
}
}
Configure BAML generator
In baml_src folder add generator.baml
generator python_client {
output_type python/pydantic
output_dir "../"
version "0.213.0"
}
The generator block tells baml-cli to create a Python client with Pydantic models in the parent directory.
When we run baml-cli generate
This will compile the .baml definitions into a baml_client/ Python package in your project root. It contains:
baml_client/types.pywith Pydantic classes (Patient, etc.).baml_client/sync_client.pyandasync_client.pywith a callablebobject. For example,b.ExtractPatientInfo(pdf)will return aPatient.
Alternative - Native ExtractByLLM Component
If you prefer to define the extraction logic in a native CocoIndex function with native Python class, you could also use the ExtractByLLM component.
You could see an example here.
Continuous Data Transformation flow with incremental processing
Next we will define data transformation flow with CocoIndex. Once you declared the state and transformation logic, CocoIndex will take care of all the state change for you from source to target.
CocoIndex Flow
Declare Flow
Declare a CocoIndex flow, connect to the source, add a data collector to collect processed data.
@cocoindex.flow_def(name="PatientIntakeExtractionBaml")
def patient_intake_extraction_flow(
flow_builder: cocoindex.FlowBuilder, data_scope: cocoindex.DataScope
) -> None:
data_scope["documents"] = flow_builder.add_source(
cocoindex.sources.LocalFile(
path=os.path.join("data", "patient_forms"), binary=True
)
)
patients_index = data_scope.add_collector()
This iterates over each document. We transform doc["content"] (the bytes) by our extract_patient_info function. The result is stored in a new field patient_info. Then we collect a row with the filename and extracted patient info.
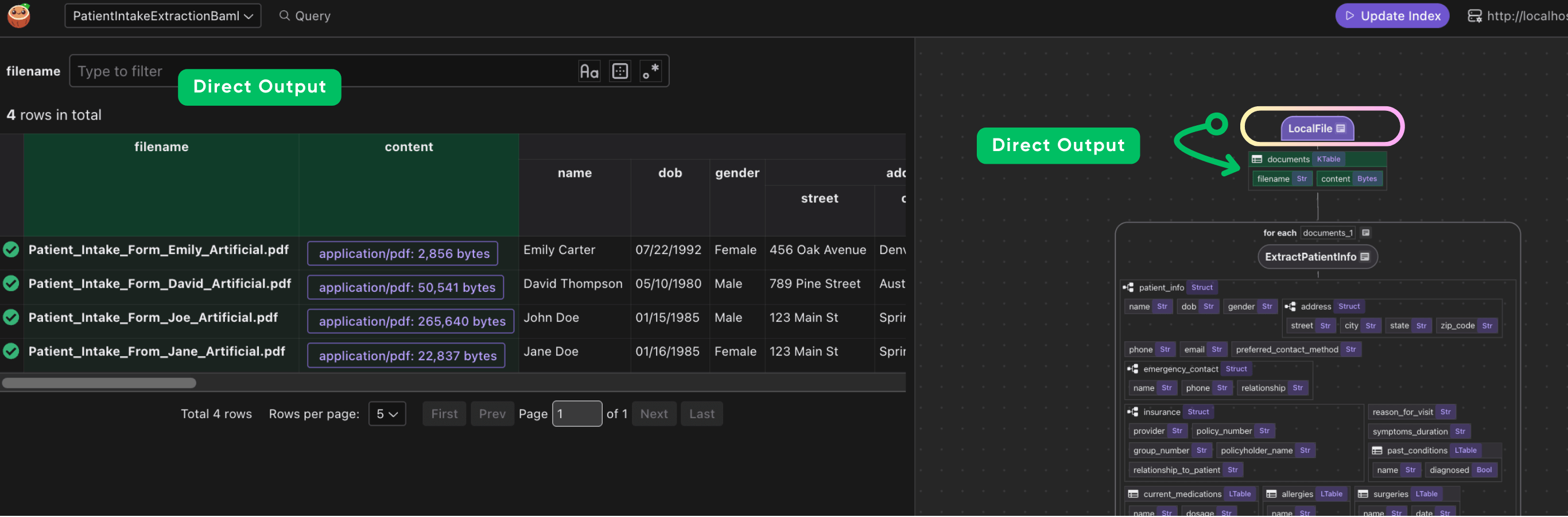
Define a custom function to use BAML extraction to transform a PDF
@cocoindex.op.function(cache=True, behavior_version=1)
async def extract_patient_info(content: bytes) -> Patient:
pdf = baml_py.Pdf.from_base64(base64.b64encode(content).decode("utf-8"))
return await b.ExtractPatientInfo(pdf)
@cocoindex.op.function(cache=True, behavior_version=1)caches results for incremental processing; bumpbehavior_versionto refresh cache if logic changes.- The function base64-encodes input bytes, creates a BAML
Pdf, and callsawait b.ExtractPatientInfo(pdf)to return aPatientobject.
Process each document
- Transform each doc with BAML
- collect the structured output
with data_scope["documents"].row() as doc:
doc["patient_info"] = doc["content"].transform(extract_patient_info)
patients_index.collect(
filename=doc["filename"],
patient_info=doc["patient_info"],
)
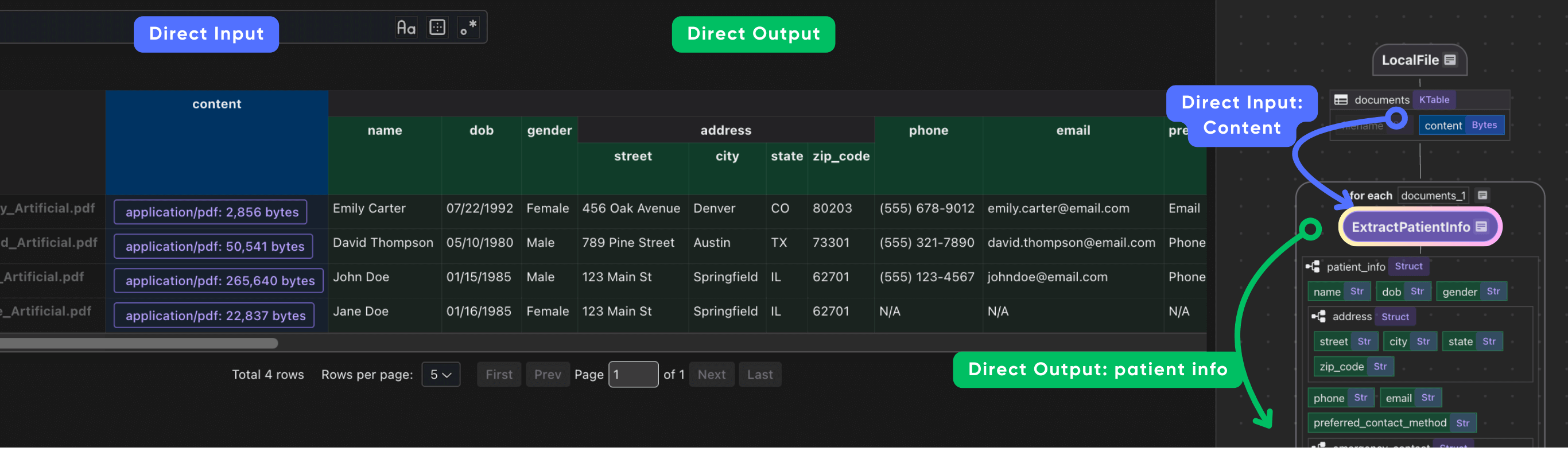
It is common to have heavy nested data, CocoIndex is natively designed to handle heavily nested data structures.
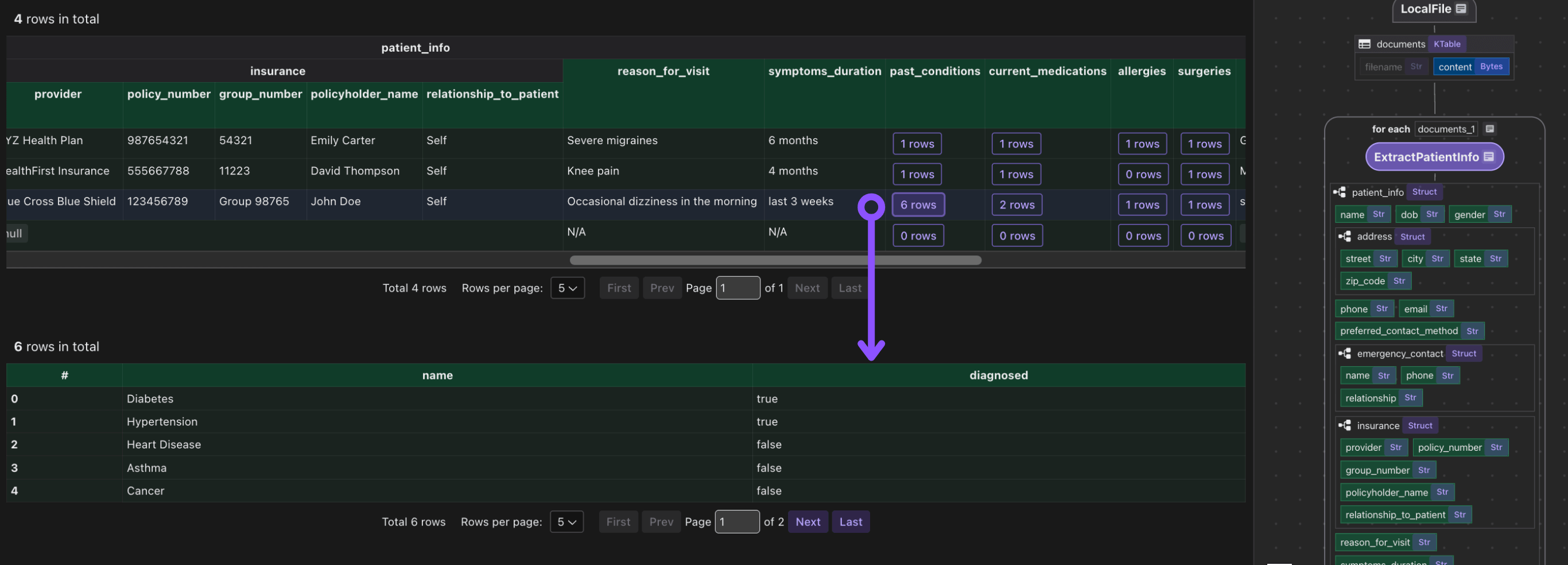
Export to Postgres
patients_index.export(
"patients",
cocoindex.storages.Postgres(),
primary_key_fields=["filename"],
)
Exports the index to Postgres as the patients table, with automatic updates and deletions when source files change.
Running the Pipeline
Generate BAML client code (required step, in case you didn’t do it earlier. )
baml generate
This generates the baml_client/ directory with Python code to call your BAML functions.
Update the index:
cocoindex update main
CocoInsight
I used CocoInsight (Free beta now) to troubleshoot the index generation and understand the data lineage of the pipeline. It just connects to your local CocoIndex server, with zero pipeline data retention.
cocoindex server -ci main
Composable by Default: Use the Best Components for Your Use Case
CocoIndex offers a comprehensive toolkit for building robust LLM pipelines, but it's intentionally built as an open and interoperable system. You can seamlessly incorporate your own preferred components—such as custom document parsers or structured extraction tools like BAML—according to your domain requirements.
Connect to other sources
CocoIndex natively supports Google Drive, Amazon S3, Azure Blob Storage, and more.
Sources